Phenotypic plasticity, genotype-by-environment interactions and genetic assimilation
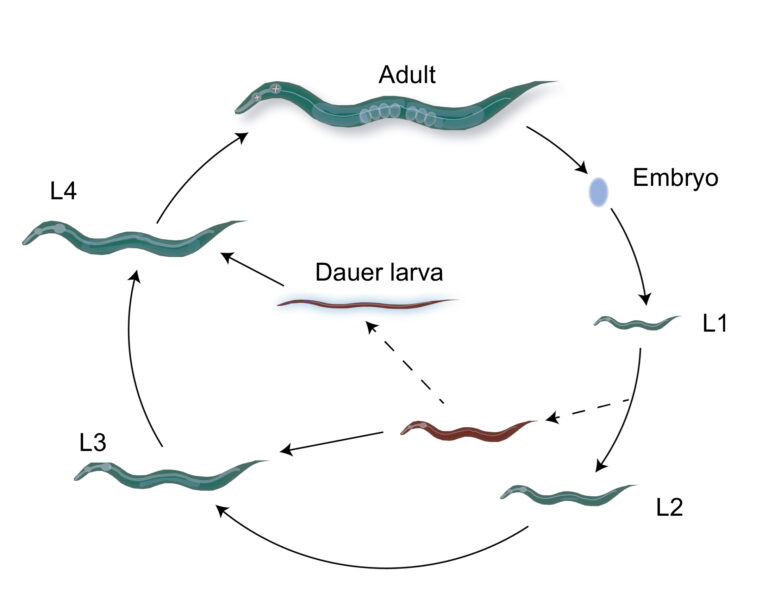
We focus on the environmental context-dependence of developmental processes (e.g. germ cell proliferation, gametogenesis, dauer formation) to characterize the molecular basis and evolution of phenotypic plasticity.